Mar 2018
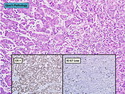
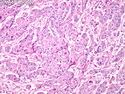
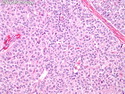
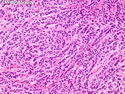
Molecular Classification of Breast Cancer
Reviewer(s): Dharam M. Ramnani, MD
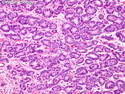
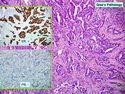
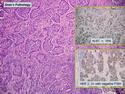
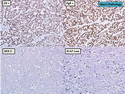
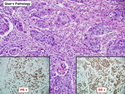
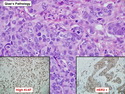
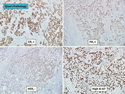
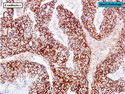
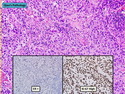
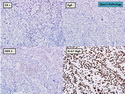
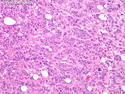
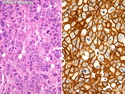
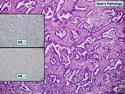
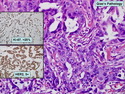
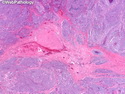
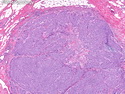
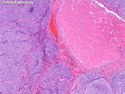
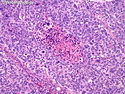
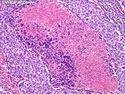
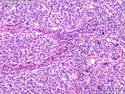
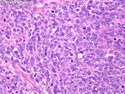
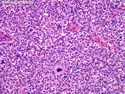
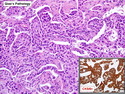
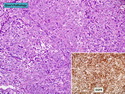
Like most other cancers, breast carcinoma exhibits an abnormal pattern of gene expression. With the advancements in technology, it has become possible to study thousands of mRNAs simultaneously from a group of tumors to develop gene expression profiles (molecular signatures). These molecular signatures may become useful for diagnosis, predicting tumor behavior, as well as selecting appropriate therapy for individual patients. The major drivers of breast cancer pathogenesis are hormone receptor-related genes, HER2 cluster of genes, and proliferation-associated genes. Based on the similarities of gene expression profiles using a hierarchical clustering, breast cancer has been subdivided into luminal A, luminal B, HER2-enriched, and basal-like subtypes. The features of individual subtypes are discussed with the photomicrographs. The clinical utility of routine molecular profiling of individual breast cancers is still uncertain. The results appear to be dependent on the platform and data analysis techniques used. Meanwhile, the traditional histopathologic grading (Nottingham combined histologic grade) is the standard of care and is unlikely to be replaced soon. References:1. DeVita, V. T., Lawrence, T. S., & Rosenberg, S. A. (2015). Cancer - Principles & Practice of Oncology: Primer of the Molecular Biology of Cancer - Second Edition. Wolters Kluwer. 2. Allison, K. H.; Molecular Pathology of Breast Cancer - What a Pathologist Needs to Know. Am J Clin Pathol 2012;138:770-780.